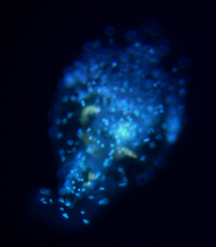
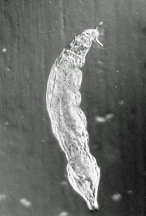
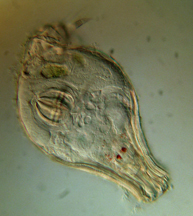
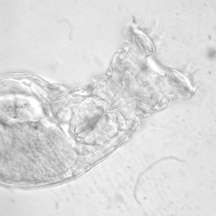
Philodina spp. from Scotia Marsh, Arizona
Current Research Projects
Mutation Accumulation in an Ancient Asexual Organism
In a mutation accumulation (MA) experiment, a
number of single asexual animals are isolated and examined daily. When
an animal reproduces, a single offspring is kept and the parent is
discarded. This is repeated for many generations. In each line, the
effective population size is 1. New mutations occur in each generation
and accumulate because there is no natural selection or random genetic
drift to eliminate them. Deleterious mutations greatly outnumber
advantageous mutations, and as they accmulate in each line the fitness
of the animals in that line decreases. One by one, they cease to
reproduce. These experiments have been applied to protists such as Paramecium aurelia and multicellular organisms such as the nematode Caenorhabditis elegans.
In our first experiment with the strictly asexual bdelloid rotifer Adineta vaga, the lines survived only 12 generations on average, and a maximum of 22 generations. In a prallel experiment Philodina roseola lines became extinct even faster. Subsequent experiments focused on A. vaga
because the complete genome of this species has been sequenced. These
experiments showed similarly rapid declines in fitness; however, we
cannot rule out the possibility that reisolation lines die because of
fitness variation unrelated to the number of detrimental mutations. To
test this, we will dehydrate and rehydrate animals and compare their
survival to that of controls in daily reisolation lines. This
dehydration-rehydration cycle is known to cause chromosomes to fragment
and then re-join, possibly with attendant repair or replacement of
detrimental alleles. Work in Claudia Ricci's lab (Universithy of Milan)
previously showed that long-term culture of bdelloids in populations of
small (unspecified, but > 1) population size caused decline in
fitness that could be reversed by a dehydration-rehydration cycle. Speciation in an Ancient Asexual
Organism
One of the major problems of biology is why most organisms
reproduce sexually at least part of the time. Theory and some experimental
evidence suggests that the loss of sexual reproduction should reduce the
effectiveness natural selection. Asexual lineages should accumulate detrimental
mutations, leading to extinction. They should also have difficulty retaining
and fixing advantageous mutations, which would make it difficult to adapt
to new environments and speciate. In fact the definition of species in asexual
organisms is controversial, since the "biological" species definition
cannot be applied. We are studying the long-term consequences of the loss
of asexual reproduction in bdelloid
rotifers, a widespread group of freshwater invertebrates which have been
reproducing asexually for at least 40 million years and undergone substantial
differentiation into species differing in morphology, habitat, and behavior.
(See the movie at BdelloidMovieShort.mov.)
We are collecting bdelloid rotifers and amplifying and
sequencing a fragment of the mitochondrial coxI gene from each isolate.
Together with Tim Barraclough and Austin Burt (Barraclough,
Birky, and Burt 2003),
we used basic population genetic theory to show
that asexual organisms, like sexual organisms, should fall into
clusters
representing independently evolving lineages. We devised a new species
concept,
the Evolutionary Genetic Species Concept, for asexuals which describes
clusters
that are comparable to biological species in sexual organisms. We also
devised
a species criterion that uses the ratio of the sequence difference
between two clades to the mean sequence difference between sequences
within one clade. This K/q
ratio, together with the number of specimens in each clade, can be used
to detemine the probability that the specimens came from two different
species. This work was described
in a preliminary paper (Birky
et al. 05)
and a more complete paper (Birky et al. 2010) where it is applied not
only to bdelloid rotifers but also to several other groups of asexual
animals and protists. With Tim Barraclough (Silwood Park), I applied the K/q
ratio and Tim's GMYC method to bdelloid rotifers and oribatid mites,
finding that the two methods agree in identifying the majority of
species from cox1 sequences (Birky and Barraclough 2009). Next I used the K/q
ratio to delimit species in an assortment of sexually-reproducing
eukaryotes (Birky 2013). Currently I am extending this to bacteria.
Note that this project is basically DNA barcoding, but the rationale
and methodology are based on well-established population genetic theory
and are thus completely different from that employed by the Barcode of
Life project.
The 2005 paper also
describes evidence that some species are adapted to different
ecological niches, which was extended to a larger sample of bdelloids
with a different method in papers from Tim's lab. Also in my 2005
paper, analyses of the same DNA sequences used for phylogenetic
analysis found that anciently asexual bdelloid rotifer lineage showed
about
the same intensity of selection (measured by Ka/Ks) as their sexual
sister
group, the monogonont rotifers.
Genetic Diversity and Sex
We have also found that the nucleotide diversity of the
cox1 gene in bdelloids is similar to that of other invertebrates,
both macroscopic and microscopic. This suggests that the effective population
size is modest, even though the census population size is immense. Evidently
bdelloids have not escaped Muller's ratchet by having extremely large effective
population sizes.
Frozen Rotifers
Bdelloid rotifers are remarkably tough. Among
other things,
they withstand dessication and disperse by blowing around in the wind
when
dessicated. Moreover, they have been found in temporary waters in
Antarctica
and on the top of 12,000-foot mountains in the U.S. Former undergrad
student
Julia Perry found that bdelloids also survive freezing at -80C, in
culture medium without cryoprotectants or any other special treatment.
We have found good survival and reproduction after freezing for more
than 2.5 years. Undergrad Alex Podolsky has investigated some of the
factors that do, or do not, affect survival after frezing at -80C.
Remarkably, they do not survive freezing at -20C, perhaps
because it is too slow and ice crystals form.
Testing the "Everything is Everywhere"
Hypothesis
We are now able to test an important hypothesis in
biogeography,
the Everything is Everywhere hypothesis. This hypothesis says that
microscopic
organisms have such large population sizes that every species can be
found
everywhere, although the local environment determines whether a species
thrives at a particular location. Some of the data favoring this
hypothesis
are based on species identified by morphology, which is often a
misleading
criterion, especially for microscopic organisms. Our preliminary data,
analyzed by EEB graduate student Erica Sommers, suggest
that although many or most bdelloid rotifer species disperse broadly
and
rapidly, not all do so and most of the species we find in the U.S. are
not
found in collections from Europe, Africa, and elsewhere. It is possible
that oceans represent significant barriers to the dispersal of
bdelloids.
Rotifer Systematics
Our collection includes a number of new species. We have
four species of Abrochtha, of which at least three are new; we collaborated with Claudia Ricci, Giulio Melone, and Diego Fontaneto of
the University of Milan to describe the new species (Birky et al. 2011). Two of them are cryptic
species, distinguishable by genotype but not be phenotype.
Link to the labs of collaborator Claudia Ricci's
web site at the University of Milan: http://users.unimi.it/ricci/rotifer.htm 
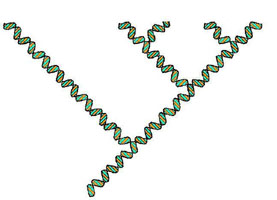
DNA barcode of Philodina roseola; each
color represents a different base in the cox1 sequence.
Link to the Barcode Blog: http://phe.rockefeller.edu/barcode/blog/
A Curious Incident
A recent paper did computer simulations which
the authors interpreted as showing that aquatic organisms smaller than
1 mm cannot form species (Rossberg et al. 2013 Proc. Roy. Soc. B
280,20131248). The authors also re-plotted data from several
experimental papers which they said supported their conclusion. Given
that many papers that have demonstrated the existence of species in
microscopic eukaryotes, including my own, and even given formal names
to some of these species, this came as a surprise! It reminded me of
the (possibly apocryphal) story of the engineer who showed
mathematically that bumblebees can't fly. A critique which describes
methodological errors in Rossberg et al. has been accepted by the Proceedings
(Morgan, Bass, Bik, Birky, et al. 2014 A critique of Rossberg et al.:
noise obscures the genetic signal of meiobiotal ecospecies in
ecogenomic datasets.). In addition to those errors, I also believe that
they have confused the large census population sizes of microscopic
organisms with the much smaller effective population sizes.
Some Representative Publications
Most of the links for downloading pdf files are broken; I'll repair them as soon as possible.
Birky, C. William, Jr., and John J. Gilbert, 1971 Parthenogenesis in
rotifers: the control of sexual and asexual reproduction. Am. Zoologist
11:245-266.
Birky, C. William, Jr.,1973 On the origin of mitochondrial mutants: Evidence
for intracellular selection of mitochondria in the origin of antibiotic-resistant
cells in yeast. Genetics 74:421-432. Birky73IntracellSelectMito.pdf
Thrailkill, Kathryn M., C. William Birky, Jr., Gudrun Lückemann,
and Klaus Wolf, 1980 Intracellular population genetics: Evidence for random
drift of mitochondrial allele frequencies in Saccharomyces cerevisiae
and Schizosaccharomyces pombe. Genetics 96:237-262. Thrailkill80MitoDrift.pdf
Birky, C. William, Jr., Karen P. VanWinkle-Swift, Barbara B. Sears, John
E. Boynton, and Nicholas W. Gillham, 1981 Frequency distributions for chloroplast
genes in Chlamydomonas zygote clones: Evidence for random drift.
Plasmid 6:173-192. Birky81ChlamyCpRandomDrift.pdf
Birky, C. William, Jr., Takeo Maruyama, and Paul Fuerst, 1983 An approach
to population and evolutionary genetic theory for genes in mitochondria
and chloroplasts, and some results. Genetics 103:513-527. Birky83OrgPopGenTheory1.pdf
Banks, Jo Ann, and C. William Birky, Jr., 1985 Chloroplast DNA diversity
is low in a wild plant, Lupinus texensis. Proc. Nat. Acad. Sci. USA
82:6950-6954. Banks85LupineCpDiversity.pdf
Birky, C. William, Jr., and J. Bruce Walsh, 1988 Effects of linkage on
rates of molecular evolution. Proc. Nat. Acad. Sci. USA 85:6414-6418.
Birky88Linkage&EvolRates.pdf
Birky, C. William, Jr., Paul Fuerst, and Takeo Maruyama, 1989 Organelle
gene diversity under migration, mutation, and drift: Equilibrium expectations,
approach to equilibrium, effects of heteroplasmic cells, and comparison
to nuclear genes. Genetics 121:613-627. Birky89OrgPopGenTheory2.pdf
Birky, C. William, Jr., 1995 Uniparental inheritance of mitochondria
and chloroplast genes: mechanisms and evolution. Proc. Nat. Acad. Sci.
USA 92:11331-11338. Birky95UPI.pdf
Rumpf, Robert, Dawne Vernon, David Schreiber, and C. William Birky, Jr.,
1996 Evolutionary consequences of the loss of photosynthesis in Chlamydomonadaceae:
Phylogenetic analysis of Rrn18 (18S rDNA) in 13 Polytoma strains
(Chlorophyta). J. Phycol. 32:119-126. Rumpf96PolytomaPhylogeny.pdf
Birky, C. William, Jr., 1996 Heterozygosity, heteromorphy, and phylogenetic
trees in asexual eukaryotes. Genetics 144:427-437. Birky96Heterozygosity.pdf
Birky, C. William, Jr., 1999 An even broader perspective on the evolution
of sex. J. Evol. Biol. 12:1013-1016. Birky99BroaderPerspective.pdf
Birky, C. William, Jr., 2001 The inheritance of genes in mitochondria
and chloroplasts: Laws, mechanisms, and models. Annu. Rev. Genet.
35:125-148. Birky01AnnRevGenet.pdf
Vernon, Dawne, Robin Gutell, Jaime Cannone, Robert Rumpf, and C. William
Birky, Jr., 2001. Accelerated evolution of functional plastid rRNA and elongation
factor genes due to reduced protein synthetic load after the loss of photosynthesis
in the chlorophyte alga Polytoma.. Mol. Biol. Evol. 18:1810-1822.
Vernon01PolyRates.pdf
Lizhi Yu, C.. William Birky, Jr., and Rodney D. Adam, 2002. The two nuclei
of Giardia each have complete copies of the genome as demonstrated
by fluorescence in situ hybridization. Eukaryotic Cell 1:191-199.
Yu02Giardia.pdf
Maughan, H., C. W. Birky,Jr, W. L. Nicholson, W. D. Rosenzweig, and R.
H. Vreeland, 2002.The paradox of the "ancient' bacterium which contains
"modern" protein-coding genes. Molecular Biology and Evolution
19:1637-1639. Maughan02BacillusPermians.pdf
Barraclough, Timothy G., C. William Birky, Jr., and Austin Burt, 2003
Diversification in sexual and asexual organisms. Evolution 57:2166-2172.
BarraBirkyBurt2003.pdf
Birky, C. William, Jr. (2004) Bdelloid rotifers revisited. Proceedings
of the National Academy of Sciences USA 101:2651-2652. Birky04BdelloidsRevisited.pdf
Birky, C. William, Jr. (2005) Sex: Is Giardia doing it in the dark?.
Current Biology 15:R56-R56. Birky05SexInGiardia?.pdf
Maughan , Heather (2004) Stochastic processes influence stationary-phase
decisions in Bacillus subtilis. Journal of Bacteriology 186:2212-2214.
Maughan04StchstcVariatn&Sel.pdf
Birky, C. William, Jr., Cynthia Wolf, Heather Maughan, Linnea Herbertson,
Elena Henry (2005) Speciation and selection without sex. Hydrobiologia
546:29-45. Birky05RotiferaX.pdf
Birky, C. William, Jr. (2008) Uniparental inheritance of organelle genes. Curr. Biol. 18:R692-R695.
Birky, C. William, Jr., Timothy G. Barraclough (2009) Asexual Speciation. In Lost Sex. The Evolutionary Biology of Parthenogenesis. Peter Van Dijk, Koen Martens, Isa Schön (eds.) Springer. pp. 201-216.
Birky, C. William, Jr. (2009). Sex and evolution in eukaryotes. in Reproduction and Developmental Biology,
edited by Andre Pires da Silva, in Encyclopedia of Life Support Systems
(EOLSS), Developed under the auspices of the UNESCO, Eolss Publishers,
Oxford, UK, [http://www.eolss.net]
Birky, C. William, Jr. (2010) Giardia sex? Yes, but how and how much? Trends Parasitol. 26:70-74.
Birky, C. William, Jr. (2010) Positively negative evidence for asexuality. J. Hered. 101(Supplement 1): 542-545.
Birky, C. William, Jr., Joshua Adams, Marlea Gemmel, Julia
Perry (2010) Using population genetic theory and DNA sequences for
species detection and identification in asexual organisms. PLoS One. 5:e10609.
Schön, Isa,
Ricardo L. Pinto, Stuart Halse, Alison J. Smith, Koen Martens & C.
William Birky, Jr. (2012) Cryptic species in putative ancient asexual
darwinulids (Crustacea, Ostracoda). PLoS One 7(7):e39844 (doi:10.1371/journal.pone.0039844).
Birky, C. William, Jr., Claudia Ricci, Giulio Melone, Diego
Fontaneto (2011) Integrating DNA and traditional taxonomy to describe
diversity in poorly studied microscopic animals: new species of the
genus Abrochtha Bryce, 1910 (Rotifera: Bdelloidea: Philodinavidae). Zool. J. Linnean Soc. 161:723-734.
Birky, C. William, Jr. (2013) Species detection and
identification in sexual organisms using population genetic theory and
DNA sequences. PLoS One 8(1):e52544.
Morgan, M.J., Bass, D., Bik, H., Birky, C.W.,
Blaxter, M., Crisp, M.D., Derycke, S., Fitch, D., Fontaneto, D., Hardy,
C.M., King, A.J., Kiontke, K.C., Moens, T., Pawlowski, J.W.,
Porazinska, D., Tang, C.Q., Thomas, W.K., Yeates, D.K., Creer, S. 2014
A critique of Rossberg et al.: noise obscures the genetic signal of
meiobiotal ecospecies in ecogenomic datasets. Proc. Roy. Soc. B 281:20133076.
Go to Research History
for a summary of Bill's previous research (this will be updated soon). Go to Publications
for a complete list of Bill's papers.
Some files are in pdf format. You can view
them with the Acrobat Reader software, available free at http://www.adobe.com.

|